In this article I will talk about a supervised machine learning tool known as Support Vector Machine (SVM). I chose to write about SVM because it is one of the most commonly used and one of the most easily-implemented machine learning techniques. In addition, its’ methodology bears some similarities to the K-Nearest Neighbor (KNN) supervised machine learning algorithm which I wrote about in my previous article.
SVM in a nutshell is a classification method. In this algorithm, we plot each observation as a point in a n-dimensional scatterplot. The “n” in this case refers to the number of classes/categories in the dataset. Although n could be bigger than 2, as the dataset in this article, the common practice is to plot 2-dimensional charts with coordinates corresponding to 2 of the classes.
To make things simple, let’s imagine we only have two classes like Height and Hair length of an individual. We would first plot these variables in a 2-dimensional space where each point has two coordinates. The coordinates are known as Support Vectors.
Let’s take a look at this example in the following scatter plot:
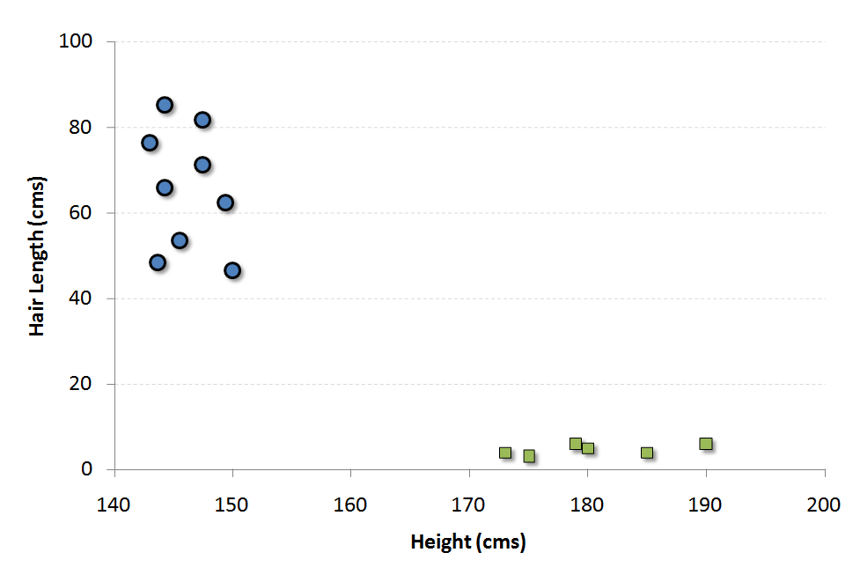
The goal of SVM is to correctly allocate all blue observations in this chart to 1 group while allocating all green observations to another group. While the allocation may be obvious in this example, we could encounter examples where points are much closer to each other than in this example. The idea of SVM could potentially solve that problem by repeatingly drawing straight lines in the chart that tries to split the data into 2 different groups. Eventually the algorithm will produce a line such that the distances from the closest point in each of the two groups are farthest away. This line is known as classifier. So whenever you add new test data into the algorithm, SVM can see where the new data lands on either side of the line, thus classifying the new data into that class. The black line in the following scatterplot illustrates the classifier line.

Now that we have a general understanding of the SVM mechanism, let’s get into the exciting part-writing R codes to test the SVM algorithm!
In R, the default package for SVM is “e1071”. So install it and call out the library on “e1071”.
> install.packages(“e1071”)
> library(e1071)
Next thing you want to do is load the “classification_spec” dataset into R. The dataset is the same as the one I used in the previous article on KNN algorithm.
> svm_data <- read.csv(“C:/Users/Shaolun/classification_spec.csv”, header=TRUE
> set.seed(1234)
Setting the seed allows you to get the same random numbers whenever you call the same seed function. By doing this you can keep track of the same random numbers easily.
> svm_split <- sample(2, nrow(svm_data), replace=TRUE, prob=c(0.67. 0.33)
The “sample” function takes a sample with a size same as the number of rows in the “svm_data” data set, which is 144, and then splits the dataset into training and testing sets. The “2” inside the function means you assign either “1” or “2” to the 144 rows of the “svm_data” dataset. The assignment of 1 has a possibility of 0.67 and assignment of 2 has possibility of 0.33. The “replace=TRUE” argument means you can assign either 1 or 2 each time of assignment.
The first 2 functions below defined the independent variables in your training and testing set, allocating the training set (1) and testing set (2) into the 4 independent variables. The 3rd and 4th functions below defines the dependent variable in training and testing set.
> svm.training <- svm_data[svm_split==1, 1:4]
> svm.testing <-svm_data[svm_split==2, 1:4]
> svm.trainLabels <- svm_data[svm_split==1, 5]
> svm.testLabels <- svm_data[svm_split==2, 5]
> summary(svm.trainLabels)
high low medium
33 37 36
> summary(svm.testLabels)
high low medium
11 15 12
Running the “summary” functions on the 2 labels allows you to verify that 106 observations are assigned to the training set and 38 assigned to testing set, which is close to the 2/3 and 1/3 ratio. Run the codes below to perform the SVM algorithm, save the result to a new variable called “fit” (or whatever variable you like to save it to as long as it’s not a reserved name in R) and see the summary. After running these codes, the “fit” variable can then be used to predict the class that a new data belongs to.
> x <- cbind(svm.training, svm.trainLabels)
> fit <-svm(svm.trainLabels ~ ., data = x)
> summary(fit)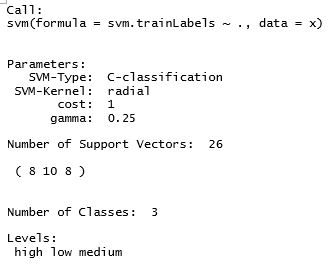
Let’s test the accuracy of SVM by predicting the class of each row in the test set.
> predicted <- predict(fit, svm.testing)
Construct data frame to compare both predicted output and actual output:
> predicted_output <- predicted
> actuals <- svm.testLabels
> data.frame(predicted, actuals)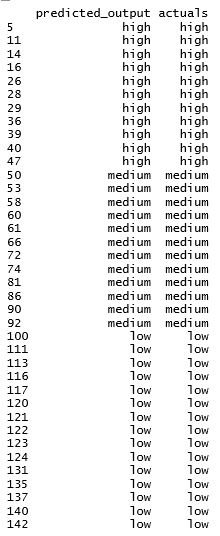
As you can see, the classifications made by SVM is 100% accurate in our example! This is very impressive considering that we only have 106 observations in the training data and we have to predict 38 observations.
To see the predicted results in scatter plot, you can run the following commands:
> predicted_output <- data.frame(svm.testing, predicted)
> qplot(GY.SiteA, GY.SiteB, data = predicted_output, color = predicted)
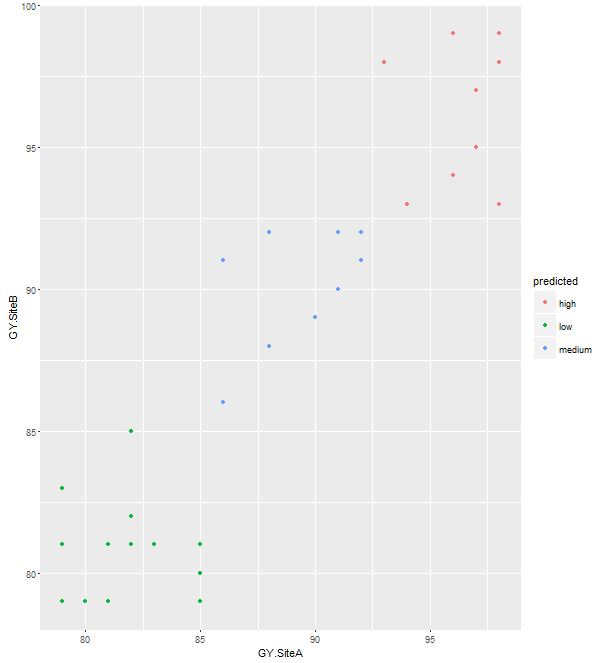
To see the plot with actual test results, run the following codes:
> actual_output <- data.frame(svm.testing, svm.testLabels)
> qplot(GY.SiteA, GY.SiteB, data = actual_output, color = svm.testLabels)
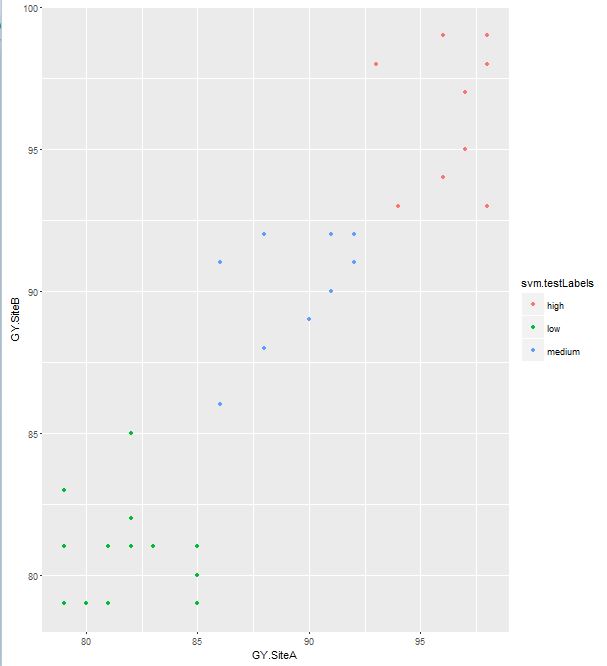
Comparing the colors in the 2 plots you can easily see that the classification of groups is exactly the same in both predicted and actual output.
So this is it for the introduction on SVM using R. Feel free to leave comments or questions in the section in the comment section below 🙂

This post is very inspiring. Also, a good idea to bridge a joint of the R programming and machine learning.
LikeLiked by 1 person
Thank you for your encouraging comments! 🙂 Like you said, R is a very powerful tool to perform many machine learning algorithms. Also important is being able to break down big data into smaller subsets so that R or Python can handle the load and be able to analyze them efficiently.
LikeLike